You signed in with another tab or window. Reload to refresh your session.You signed out in another tab or window. Reload to refresh your session.You switched accounts on another tab or window. Reload to refresh your session.Dismiss alert
This may be a followup of a previous closed issue (#118).
I am making polar stereographic projection pcolormesh plots of some sea ice data. It works fine with the Northern Hemisphere, but produces solid color for the South.
The problem was replicated on both Linux and Windows with different versions of Python, numpy, matplotlib and basemap. I am using basemap-1.1.0 with matplotlib-1.5.3 in winpython-3.5 on Windows 10 to produced the following figures.
Here is an example script:
from netCDF4 import Dataset
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import array
import matplotlib.cm as cm
from mpl_toolkits.basemap import Basemap
import glob
import struct
import datetime
import time
import sys
from pylab import *
POLE=sys.argv[3]
fname=sys.argv[1]
fld=sys.argv[2]
print(fname)
ncfile = Dataset(fname, 'r', format='NETCDF4')
fbot=ncfile.variables[fld][0]
if 'lon' in ncfile.variables:
LON=ncfile.variables['lon'][:]
if 'lon_scaler' in ncfile.variables:
LON=ncfile.variables['lon_scaler'][:]
if 'LON' in ncfile.variables:
LON=ncfile.variables['LON'][:]
if 'lat' in ncfile.variables:
LAT=ncfile.variables['lat'][:]
if 'lat_scaler' in ncfile.variables:
LAT=ncfile.variables['lat_scaler'][:]
if 'LAT' in ncfile.variables:
LAT=ncfile.variables['LAT'][:]
if 'kmt' in ncfile.variables:
kmt=ncfile.variables['kmt'][:]
lon=LON
lat=LAT
if len(LON.shape) == 1:
lon,lat=np.meshgrid(LON,LAT)
if 'kmt' in ncfile.variables:
fbot=ma.masked_where(kmt<1, fbot)
ncfile.close()
meridians=[1,0,1,1]
fig=figure(figsize=(12,12), facecolor='w')
if POLE=='N':
m = Basemap(projection='npstere',lon_0=0,boundinglat=45)
if POLE=='S':
m = Basemap(projection='spstere',lon_0=180,boundinglat=-45)
m.drawcoastlines()
m.fillcontinents()
m.drawcountries()
plt.title(fld)
x, y =m(lon,lat)
m.pcolormesh(x,y,fbot,vmin=float(sys.argv[4]), vmax=float(sys.argv[5]))
m.drawparallels(np.arange(-90.,120.,15.),labels=[1,0,0,0]) # draw parallels
m.drawmeridians(np.arange(0.,420.,30.),labels=meridians) # draw meridians
plt.colorbar(orientation='vertical',extend='both',shrink=0.8)
plt.show()
A netcdf file to reproduce the problem can be downloaded here
Hi,
This may be a followup of a previous closed issue (#118).
I am making polar stereographic projection pcolormesh plots of some sea ice data. It works fine with the Northern Hemisphere, but produces solid color for the South.
The problem was replicated on both Linux and Windows with different versions of Python, numpy, matplotlib and basemap. I am using basemap-1.1.0 with matplotlib-1.5.3 in winpython-3.5 on Windows 10 to produced the following figures.
Here is an example script:
A netcdf file to reproduce the problem can be downloaded here
ftp://pscftp.apl.washington.edu/zhang/Global_seaice/heff.H1987.nc.gz
Once untared, the following call to the above script (named plot_field_pole.py)
python plot_field_pole.py heff.H1987.nc heff S 0.0 3.0
will give the wrong plot.
while
python plot_field_pole.py heff.H1987.nc heff N 0.0 3.0
produced correct one.
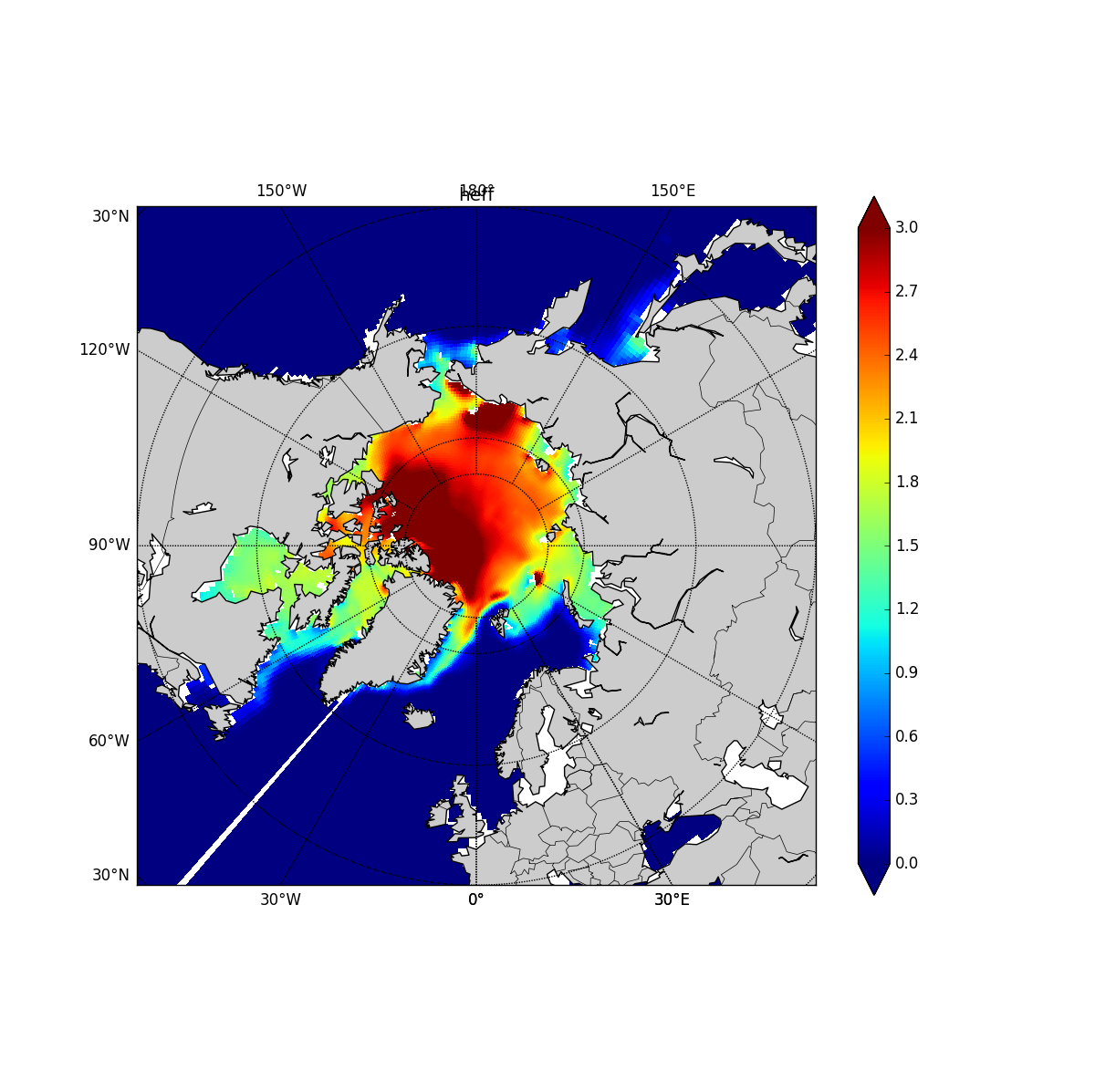
The only difference between the two commands is Basemap(projection='spstere') or
Basemap(projection='npstere')
Any help will be much appreciated.
Bin Zhao
The text was updated successfully, but these errors were encountered: